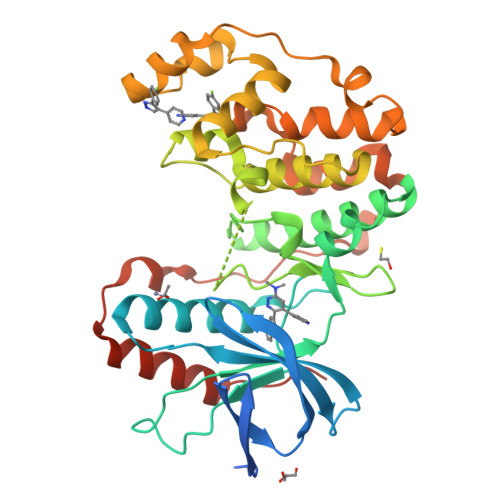
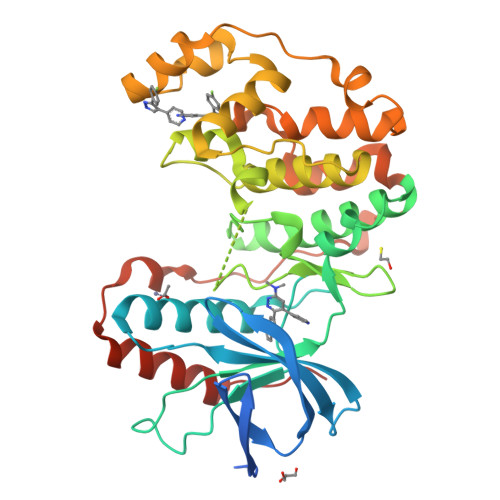
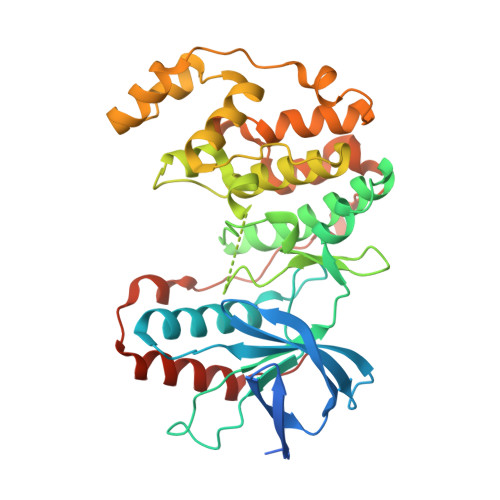
Development of Novel In Vivo Chemical Probes to Address CNS Protein Kinase Involvement in Synaptic Dysfunction.
Watterson, D.M., Grum-Tokars, V.L., Roy, S.M., Schavocky, J.P., Bradaric, B.D., Bachstetter, A.D., Xing, B., Dimayuga, E., Saeed, F., Zhang, H., Staniszewski, A., Pelletier, J.C., Minasov, G., Anderson, W.F., Arancio, O., Van Eldik, L.J.(2013) PLoS One 8: e66226-e66226
- PubMed: 23840427
- DOI: https://doi.org/10.1371/journal.pone.0066226
- Primary Citation of Related Structures:
4F9W, 4F9Y, 4FA2 - PubMed Abstract:
Serine-threonine protein kinases are critical to CNS function, yet there is a dearth of highly selective, CNS-active kinase inhibitors for in vivo investigations. Further, prevailing assumptions raise concerns about whether single kinase inhibitors can show in vivo efficacy for CNS pathologies, and debates over viable approaches to the development of safe and efficacious kinase inhibitors are unsettled. It is critical, therefore, that these scientific challenges be addressed in order to test hypotheses about protein kinases in neuropathology progression and the potential for in vivo modulation of their catalytic activity. Identification of molecular targets whose in vivo modulation can attenuate synaptic dysfunction would provide a foundation for future disease-modifying therapeutic development as well as insight into cellular mechanisms. Clinical and preclinical studies suggest a critical link between synaptic dysfunction in neurodegenerative disorders and the activation of p38αMAPK mediated signaling cascades. Activation in both neurons and glia also offers the unusual potential to generate enhanced responses through targeting a single kinase in two distinct cell types involved in pathology progression. However, target validation has been limited by lack of highly selective inhibitors amenable to in vivo use in the CNS. Therefore, we employed high-resolution co-crystallography and pharmacoinformatics to design and develop a novel synthetic, active site targeted, CNS-active, p38αMAPK inhibitor (MW108). Selectivity was demonstrated by large-scale kinome screens, functional GPCR agonist and antagonist analyses of off-target potential, and evaluation of cellular target engagement. In vitro and in vivo assays demonstrated that MW108 ameliorates beta-amyloid induced synaptic and cognitive dysfunction. A serendipitous discovery during co-crystallographic analyses revised prevailing models about active site targeting of inhibitors, providing insights that will facilitate future kinase inhibitor design. Overall, our studies deliver highly selective in vivo probes appropriate for CNS investigations and demonstrate that modulation of p38αMAPK activity can attenuate synaptic dysfunction.
Organizational Affiliation:
Department of Molecular Pharmacology and Biological Chemistry, Northwestern University, Chicago, Illinois, United States of America.